This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Protein Alignment and Phylogeny
The NOD2 protein sequences from all 8 species were aligned using multiple sequence alignment tools availible through the European Bioinformatics Institute. Below are the sequence alignments using ClustalW2, MUSCLE, and T-Coffee.
|
|
|
||||||||||||||||||
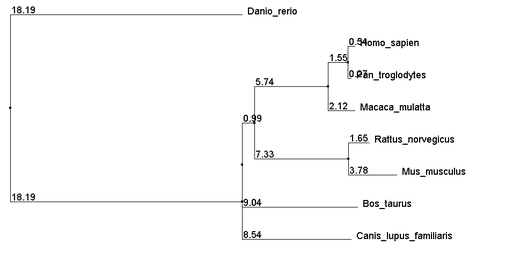
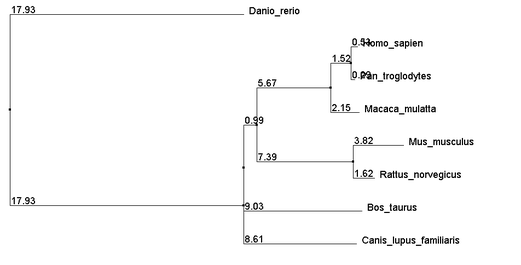
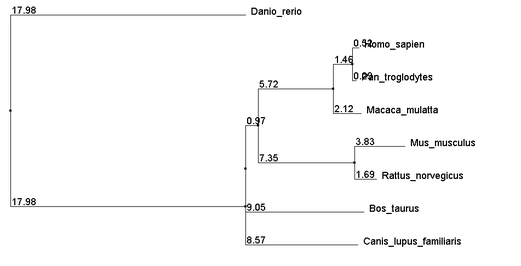
The Above alignments were then used to build neighbor joining trees using percent identity. The resulting trees are as follows:
Analysis
The close relations between Human and Chimpanzee as well as between Mouse and Rat agree with the TreeFam gene map seen at the bottom of the gene phylogeny page. Note that trees built from each of the three alignment methods yield identical topologies. The distances also show very close agreement. Given that all 3 methods of alignment generated very similar trees, it is quite likely that the relationships suggested by the trees are highly accurate.
It is unlikely, however, that the polytomy containing Dog, Cow, and the Human-Rat clade is a real polytomy, but additional analysis was unable to resolve it. From these trees we get more support for the strength mice as a model organism for the disease.