This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Conclusions
Throughout the various pages of this site, we have explored NOD2 and its role in Crohn's Disease(CD). A review of the literature as well as our genomic and proteomic analyses of NOD2 have provided great insight into NOD2's role in CD. There remains much to be learned, however. One of the greatest issues left to be resolved is exactly how NOD2 mutation leads to the increased inflammation associated with CD. While most variants associated with the disease inhibit the protein's ability to recognize and respond to bacterial infection, the end result is over-activation, rather than inactivation of the immune response. Currently the two leading schools of thought are 1. that faulty NOD2 allows initial bacterial infection to take place, at which point parallel immune systems over-respond and 2. that NOD2 is also involved in negative regulation of immune response, and faulty NOD2 is no longer able to take place in said regulation, leading to over-response. This is no small task, and it will involve a more extensive understanding of what other proteins interact with NOD2 than is currently known. In any case, NOD2's role in CD is clear based on Genome Wide Association Study (GWAS) as well as monozygotic twin studies [1].
Below is a link to a powerpoint presentation outlining proposed research into NOD2 interactions. The proposal is also summarized in text below.
Below is a link to a powerpoint presentation outlining proposed research into NOD2 interactions. The proposal is also summarized in text below.
| NOD2 and Crohn's Disease Presentation | |
| File Size: | 3370 kb |
| File Type: | pptx |
Research
Given the controversy remaining as to by which mechanisms faulty NOD2 leads to CD, I chose to focus my research directly on illuminating and exploring novel NOD2 protein-protein interactions. The search began with the discovery of NLRP1 and NLRC4(CLAN) in my probe of protein interaction networks. Both NLRP1 and CLAN have been shown to bind directly to NOD2, forming hetero-oligomers, suggesting the possibility that NOD2 may form such oligomers with other proteins common to GI immune response [6,7]. My main hypothesis is that NOD2 forms hetero-oligomers by NACHT domain interaction with many similar proteins. These hetero-oligomers then regulate downstream immune response via the activity of their CARD domains. Given NOD2's normal response to bacterial muramyl dipeptide (MDP), it is reasonable to expect different interactions in the presence and absence of this compound. Identification of proteins interacting with NOD2 in this way will provide novel targets for drug treatment of CD and will improve our understanding of CD pathogenesis.
http://smart.embl.de/
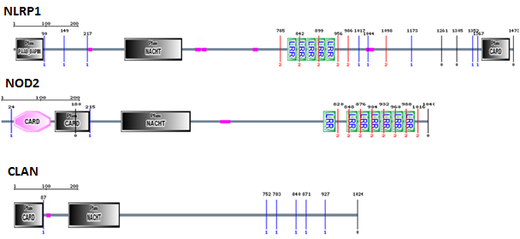
To begin, I used SMART to look at what NOD2's known binding partners, NLRP1 and CLAN, had in common. NLRP1, NOD2, and CLAN all contain NACHT domains crucial for oligomerization as well as CARD domains required for activating downstream pathways once oligomerization takes place. NOD2 and NLRP1 also have LRR domains, critical for their recognition of MDP required for their activation. Based on this, I decided to probe the online databases for proteins which had NACHT and LRR domains similar to those of NOD2. I blasted the entire NACHT domain as well as the C-terminal 100 amino acids of the LRR domain, based on the fact that truncation analysis has shown that MDP recognition requires this terminal 100 amino acid region [3].
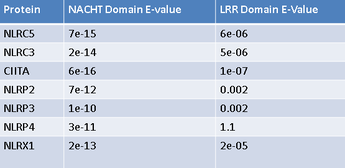
Based on this, I chose to focus my efforts on proteins with similar NACHT and LRR domains. I BLASTed both said NOD2 domains and cross referenced the lists generated for each domain to find proteins with high similarity in both domains. To the left is a list of the top human candidates for NOD2 interacting proteins based on this method. Before I turned my efforts towards testing these potential interactions I needed to establish a model in which to work. Based on protein homology, phylogeny, and interaction networks I chose to work in an human cell culture system. Though a transgenic mouse model would be a good resourcel based on phylogeny and homology, NLRP1-NOD2 and CLAN-NOD2 interactions have not been shown in mice. Additionally, much NOD2 research has been done using human tissue culture, specifically NCM460 cells [4]. This cell line is generated from healthy epithelial colon cells, so is expected to be expressing NOD2 and all CD relevant proteins at normal levels, providing a great model in which to work.
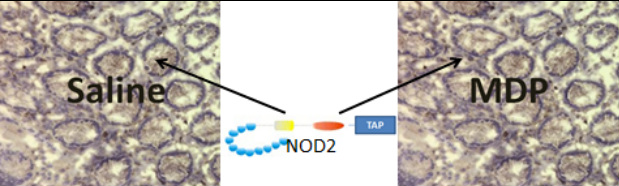
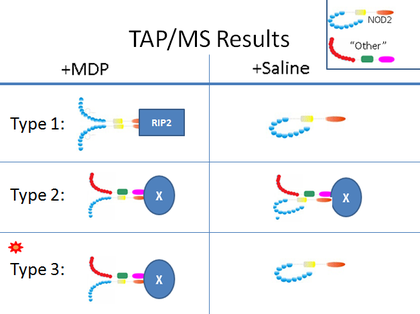
In order to test for interaction, I plan to use TAP pull down of TAP-tagged NOD2 followed by mass spectrometry for identification of associating proteins. Given the importance of the C-terminal LRR domains in MDP recognition, I will use N-terminal TAP-tagging of NOD2 expressed in NCM460 cells. In order to test for varied expression in response to bacterial MDP, I will culture two groups of cells, one in high concentration MDP solution and one control line raised in saline. I expect to find the above listed proteins, as well as new proteins, in association with NOD2. I also expect to find different binding partnerships in the presence and absence of MDP. Below is a table of the three main interactions I expect to find.
|
Type 1 represents currently known NOD2 function. NOD2 homo-oligomerizes in the presence of MDP and then initiates immune response by activating RIP2 kinase with its CARD domians.
Type 2 represents hetero-oligomers which are active regardless of MDP presence. Here the "X" represents the target of CARD domain activation. Type 3 represents hetero-oligomers which form and activate downstream response through their CARD domains only in the presence of MDP. This is the most interesting group in regards to CD because mutant NOD2 which fails to recognize MDP would also disrupt these interactions. The next step will be to identify which systems the associated "X" proteins are involved with. |
Future Directions
Once we have a group of these type 3 "X" proteins we will use additional TAP pull downs to determine in which pathway they play a role. Also we will use microarray data from CD patients and controls to identify the "X" proteins who have altered activity in CD. Once we have identified the proteins with altered function in CD, we can begin to target them as sites for therapy. A chemical screen should be performed on each of these proteins to find molecules which increase and/or inhibit their function. Hopefully compounds will be identified which will allow CD treatment directly at the level of MDP recognition, rather than simply treating the downstream inflammatory symptoms.
References
[1] Nakagome S, Mano S, Kozlowski L, et al. Crohn's Disease Risk Alleles on the NOD2 Locus Have Been Maintained by Natural Selection on Standing Variation. Mol Biol Evol. 2012;29(6):1569-85.
[2] SMART
[3] Tanabe T, Chamaillard M, Ogura Y, et al. Regulatory regions and critical residues of NOD2 involved in muramyl dipeptide recognition. EMBO J. 2004;23(7):1587-97.
[4 ] Zeng H, Trujillo ON, Moyer MP, Botnen JH. Prolonged sulforaphane treatment activates survival signaling in nontumorigenic NCM460 colon cells but apoptotic signaling in tumorigenic HCT116 colon cells. Nutr Cancer. 2011;63(2):248-55.
[5] Borthakur A, Bhattacharyya S, Dudeja PK, Tobacman JK. Carrageenan induces interleukin-8 production through distinct Bcl10 pathway in normal human colonic epithelial cells. Am J Physiol Gastrointest Liver Physiol. 2007;292(3):G829-38.
[6] Damiano JS, Oliveira V, Welsh K, Reed JC. Heterotypic interactions among NACHT domains: implications for regulation of innate immune responses. Biochem J. 2004;381(Pt 1):213-9.
[7] Hsu, L.-C., S. R. Ali, S. McGillivray, P.-H. Tseng, S. Mariathasan, E. W. Humke, L. Eckmann, J. J. Powell, V. Nizet, V. M. Dixit, and M. Karin. "A NOD2-NALP1 Complex Mediates Caspase-1-dependent IL-1 Secretion in Response to Bacillus Anthracis Infection and Muramyl Dipeptide." Proceedings of the National Academy of Sciences 105.22 (2008): 7803-808. Print. .
[2] SMART
[3] Tanabe T, Chamaillard M, Ogura Y, et al. Regulatory regions and critical residues of NOD2 involved in muramyl dipeptide recognition. EMBO J. 2004;23(7):1587-97.
[4 ] Zeng H, Trujillo ON, Moyer MP, Botnen JH. Prolonged sulforaphane treatment activates survival signaling in nontumorigenic NCM460 colon cells but apoptotic signaling in tumorigenic HCT116 colon cells. Nutr Cancer. 2011;63(2):248-55.
[5] Borthakur A, Bhattacharyya S, Dudeja PK, Tobacman JK. Carrageenan induces interleukin-8 production through distinct Bcl10 pathway in normal human colonic epithelial cells. Am J Physiol Gastrointest Liver Physiol. 2007;292(3):G829-38.
[6] Damiano JS, Oliveira V, Welsh K, Reed JC. Heterotypic interactions among NACHT domains: implications for regulation of innate immune responses. Biochem J. 2004;381(Pt 1):213-9.
[7] Hsu, L.-C., S. R. Ali, S. McGillivray, P.-H. Tseng, S. Mariathasan, E. W. Humke, L. Eckmann, J. J. Powell, V. Nizet, V. M. Dixit, and M. Karin. "A NOD2-NALP1 Complex Mediates Caspase-1-dependent IL-1 Secretion in Response to Bacillus Anthracis Infection and Muramyl Dipeptide." Proceedings of the National Academy of Sciences 105.22 (2008): 7803-808. Print. .