This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Gene Ontology
Gene ontology aims to form a database of gene products using a standardized vocabulary. This process compiles ontologies for gene products starting with three basic catagories: cellular components, molecular function, and biological process [2]. Below are lists of pertinent ontology information for human NOD2 acquired using AmiGO.
Cellular Components:
Molecular Function:
ATP binding
CARD domain binding
enzyme binding
muramyl dipeptide binding peptidoglycan binding protein binding protein kinase binding
CARD domain binding
enzyme binding
muramyl dipeptide binding peptidoglycan binding protein binding protein kinase binding
Biological Process:
defense response to bacterium defense response to Gram-positive bacterium detection of bacterium
detection of muramyl dipeptide
innate immune response in mucosa
intracellular signal transduction
positive regulation of NF-kappaB transcription factor activity
positive regulation of type 2 immune response
protein oligomerization
response to muramyl dipeptide
Toll signaling pathway
detection of muramyl dipeptide
innate immune response in mucosa
intracellular signal transduction
positive regulation of NF-kappaB transcription factor activity
positive regulation of type 2 immune response
protein oligomerization
response to muramyl dipeptide
Toll signaling pathway
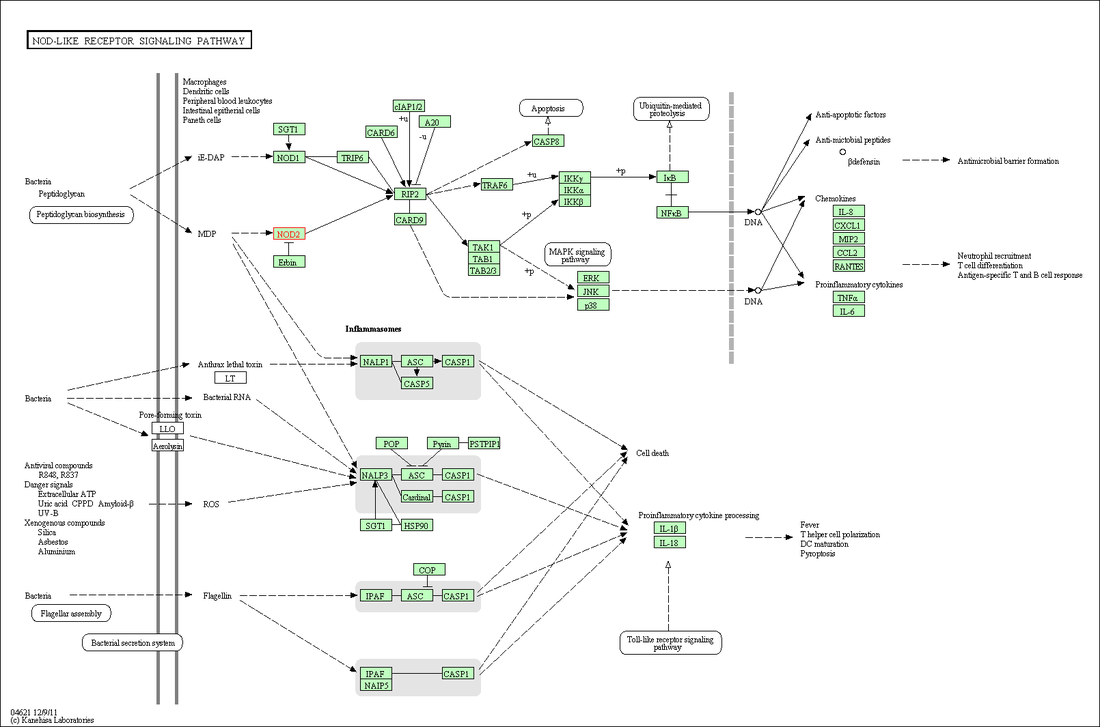
NOD2 Pathway
Analysis
Analysis of the ontology terms for NOD2 provided by
AmiGO show how it is involved in regulation of inflammatory immune response. NOD2 appears to be located throughout the entire cell, with the one exception being the nucleus. Outside of the obvious immune functions, biological functions and molecular processes involving muramyl dipeptide(MDP) came up several times in AmiGO analysis, suggesting its importance in NOD2 activity. Above is a proposed pathway for MDP initiation and downstream inflammatory response taken from KEGG [1]. The involvement in in a Toll-like signalling pathway also suggests that NOD2 is involved in regulating more than just RIP2 induced immune response, providing other avenues in which to pursue research. For example, it has been suggested that, though NOD2 is involved in upregulation of NFkB activity through RIP2, NOD2 may also inhibit NFkB through a toll-like signalling pathway. Thus a mutant NOD2, unable to interact in the toll-like signalling pathway, may result in less repression and thus more activation of NFkB [3]. Understanding the balance between this toll-like signalling pathway and the RIP2 pathway will be critical critical in understanding Crohn's Disease pathogenesis.