This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
DNA motifs
MOTIF was used to search the PROSITE database to identify the following 14 motifs in the full NOD2 gene :
|
Motif: EGF_1
PrositeID: PS00022 Description: EGF-like domain signature 1 Pattern: C-x-C-x(2)-{V}-x(2)-G-{C}-x-C Information: Commonly found in extracellular domain of membrane proteins Motif: J_ACTX PrositeID: PS60020 Description: Janus-faced atracotoxin (J-ACTX) family signature. Pattern: C-{C}(6)-C-x(2)-C-C-x-C-C-x(4)-C-x(9,10)-C Information: class of insecticidal neurotoxins Motif: INTEGRIN_BETA PrositeID: PS00243 Description : Integrins beta chain cysteine-rich domain signature Pattern: C-x-[GNQ]-x(1,3)-G-x-C-x-C-x(2)-C-x-C Information: cell surface receptors involved in cell-cell adhesion Motif: CTCK_1 PrositeID: PS01185 Description : C-terminal cystine knot signature Pattern: C-C-x(13)-C-x(2)-[GN]-x(12)-C-x-C-x(2,4)-C Information: a twisted pair of beta-strands Motif: ANAPHYLATOXIN_1 PrositeID: PS01177 Description: Anaphylatoxin domain signature Pattern: [CSH]-C-x(2)-[GAP]-x(7,8)-[GASTDEQR]-C-[GASTDEQL]-x(3,9)-[GASTDEQN]-x(2)- [CE]-x(6,7)-C-C Information: mediate inflammation through smooth muscle contraction Motif: I_CONOTOXIN PrositeID : PS60019 Description : I-superfamily conotoxin signature. Pattern: C-{C}(6)-C-{C}(5)-C-C-x(1,3)-C-C-x(2,4)-C-x(3,10)-C Information: small, neurotoxic peptides Motif: DEFENSIN PrositeID: PS00269 Description: Mammalian defensins signature Pattern: C-x-C-x(3,5)-C-x(7)-G-x-C-x(9)-C-C Information: antibacterial peptides |
Motif: IGFBP_N_1
PrositeID: PS00222 Description: Insulin-like growth factor-binding protein (IGFBP) N-terminal domain signature Pattern: GP]-C-[GSET]-[CE]-[CA]-x(2)-C-[ALP]-x(6)-C Information: bind specific extracellular proteins Motif: THIOLASE_3 PrositeID: PS00099 Description: Thiolases active site Pattern: [AG]-[LIVMA]-[STAGCLIVM]-[STAG]-[LIVMA]-C-{Q}-[AG]-x-[AG]-x-[AG]-x-[SAG] Information: enzymes involved in metabolism Motif: TUBULIN PrositeID: PS00227 Description: Tubulin subunits alpha, beta, and gamma signature Pattern: [SAG]-G-G-T-G-[SA]-G Information: principle component of microtubules Motif: HYDROPHOBIN PrositeID: PS00956 Description: Fungal hydrophobins signature Pattern: [GN]-[DNQPSA]-x-C-[GSTANK]-[GSTADNQ]-[STNQI]-[PTIV]-x-C-C-[DENQKPST] Information: hydrophobic proteins common to fungal spores Motif: VWFC_1 PrositeID: PS01208 Description : VWFC domain signature Pattern: C-x(2,3)-C-{CG}-C-x(6,14)-C-x(3,4)-C-x(2,10)-C-x(9,16)-C-C-x(2,4)-C Information: common to blood plasma proteins Motif: 4FE4S_FER_1 PrositeID: PS00198 Description : 4Fe-4S ferredoxin-type iron-sulfur binding region signature Pattern: C-x-{P}-C-{C}-x-C-{CP}-x-{C}-C-[PEG] Information: involved in metabolic electron transfer Motif: 2FE2S_FER_1 PrositeID: PS00197 Description: 2Fe-2S ferredoxin-type iron-sulfur binding region signature Pattern: C-{C}-{C}-[GA]-{C}-C-[GAST]-{CPDEKRHFYW}-C Information: involved in metabolic electron transfer |
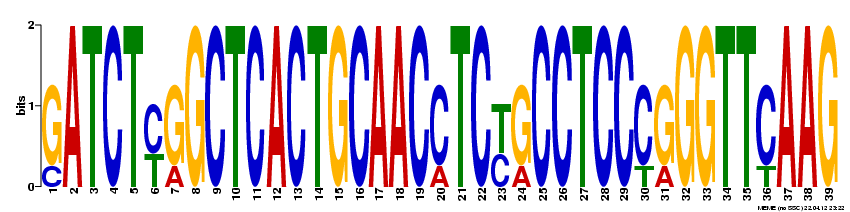
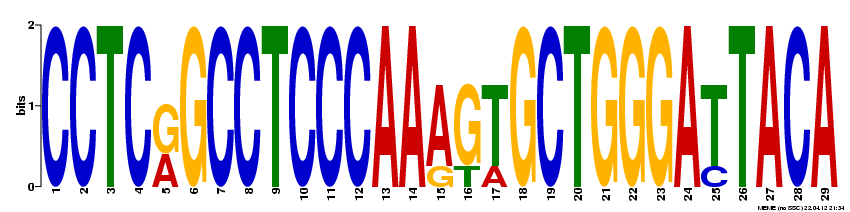
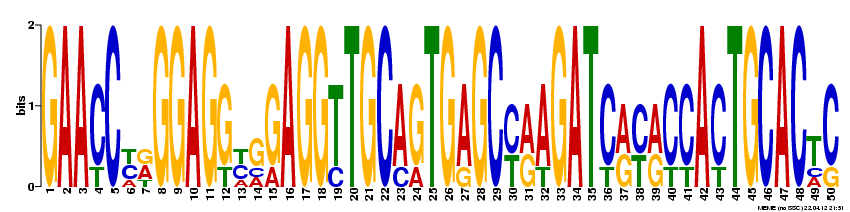
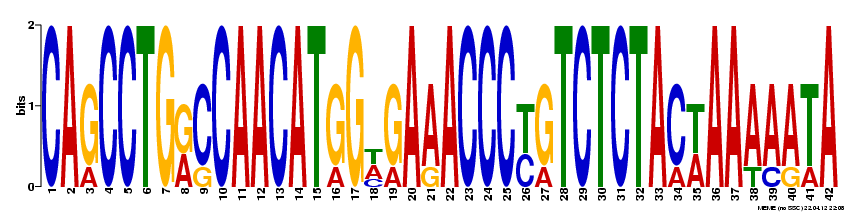
MEME was also used to probe the full NOD2 gene for motifs likely to serve a biological function. The following four motifs were detected:
Analysis:
While the potential motifs identified by MEME may prove useful jumping off points for future work, the results obtained using MOTIF directly aid our current understanding of NOD2 function. The
DEFENSIN,
ANAPHYLATOXIN_1, and
IGFBP_N_1 motifs agree well with our understanding of NOD2's role in bacterial response and inflammation. The presence of
Janus-faced atracotoxin (J-ACTX) family signature as well as I-superfamily conotoxin signature were surprising, as they are mainly found in spider venom and cone snail venom, respectively. The presence of these toxin motifs is potentially due to NOD2's role in digestion. It has been shown that many venoms have high similarity to digestive enzymes, suggesting a possible explanation for the origin of such venoms as a survival tactic [5]. Though this may not provide much insight into Crohn's disease, it is an interesting piece of evidence for the evolutionary origins of venoms.
References
[1] MOTIF
[2] PROSITE
[3] MEME
[4] Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994.
[5] "BioQuick News--Life Science News from Around the Globe." Convergent Evolution of Toxins Seen in Shrew and Lizard. Web. <http://www.bioquicknews.com/node/204>.
[2] PROSITE
[3] MEME
[4] Timothy L. Bailey and Charles Elkan, "Fitting a mixture model by expectation maximization to discover motifs in biopolymers", Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology, pp. 28-36, AAAI Press, Menlo Park, California, 1994.
[5] "BioQuick News--Life Science News from Around the Globe." Convergent Evolution of Toxins Seen in Shrew and Lizard. Web. <http://www.bioquicknews.com/node/204>.