This web page was produced as an assignment for Genetics 677, an undergraduate course at UW-Madison.
Gene Alignment and Phylogeny
The human NOD2 gene and its homologs are long enough to make sequence comparison and alignment impossible using MUSCLE and T-Coffee. Also note that the sequencs for Dog and Rhesus Macaque are predictions of mRNA sequence and thus represent cDNA. This limits direct comparison to full genomic data. Nevertheless, the NOD2 gene sequences from all 8 homologs were aligned using ClustalW2.
| ClustalW2 Gene Alignment | |
| File Size: | 668 kb |
| File Type: | clustalw |
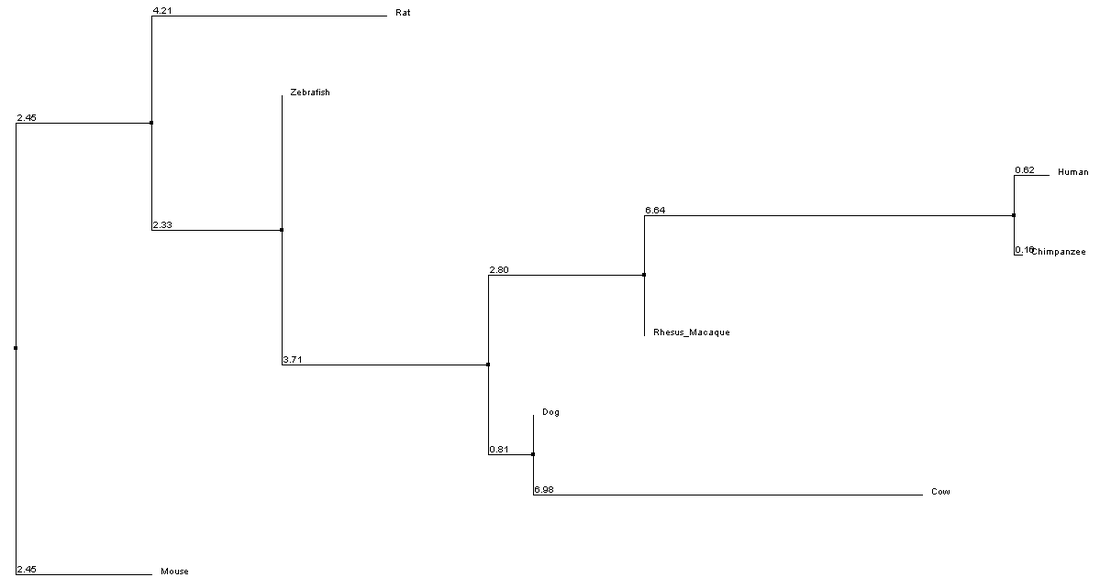
The above ClustalW2 alignment was used to build the following neighbor joining tree using percent identity.
Analysis
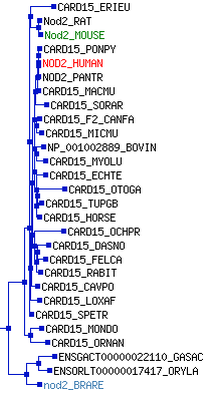
The above tree disagrees with the trees generated from protein alignments. Primarily, mouse and rat are more distantly related to human in this genetic tree than in the protein trees. This can likely be explained by poor alignment due to the increased length of the mouse and rat genes. TreeFam was used to verify that mouse and rat NOD2 are in fact more closely related to Human NOD2 than zebrafish. The Image to the right shows a gene sequence-based phylogeny taken from TreeFam. Note that Mouse(in green) and Rat Nod2 are very closely related. Likewise note that Human (in red) and Chimpanzee(PANTR) NOD2 share more with one another than the rest of the tree. The Zebrafish version of nod2 (blue) is the most distantly related of all homologs in TreeFam, agreeing well with the protein alignments seen here.